Purdue researchers use quantum-biochemical calculations to show key SARS-CoV-1 coronavirus behavior
2021-01-13
Writer(s): Cheryl Pierce
This discovery could potentially guide future development of antiviral drugs
Coronaviruses have been widely recognized as adaptive and can quickly infect large populations. Most recently SARS-C0V-2, the coronavirus that induces COVID-19 disease, has spread throughout the world and infected millions. Back in 2002, there was an outbreak produced by another coronavirus, known as SARS-CoV-1, which rapidly spread throughout several countries. What these coronaviruses have in common is a roughly spherical shape with outer spikes, called spike (S) proteins, which interact with human cells to initiate the infection. It is critical to learn what might attract or detract this initial connection with the human cells so that antiviral drugs or vaccines can be developed to prevent these interactions.
Dr. Jorge H. Rodríguez, Associate Professor of Physics and Astronomy at Purdue University, and Akshita Gupta, graduate student, have recently published a paper in Scientific Reports titled “Contact residue contributions to interaction energies between SARS-CoV-1 spike proteins and human ACE2 receptors” which identifies, using computational quantum-biochemical calculations, which viral fragments are attractive and which are repulsive to the human cells.
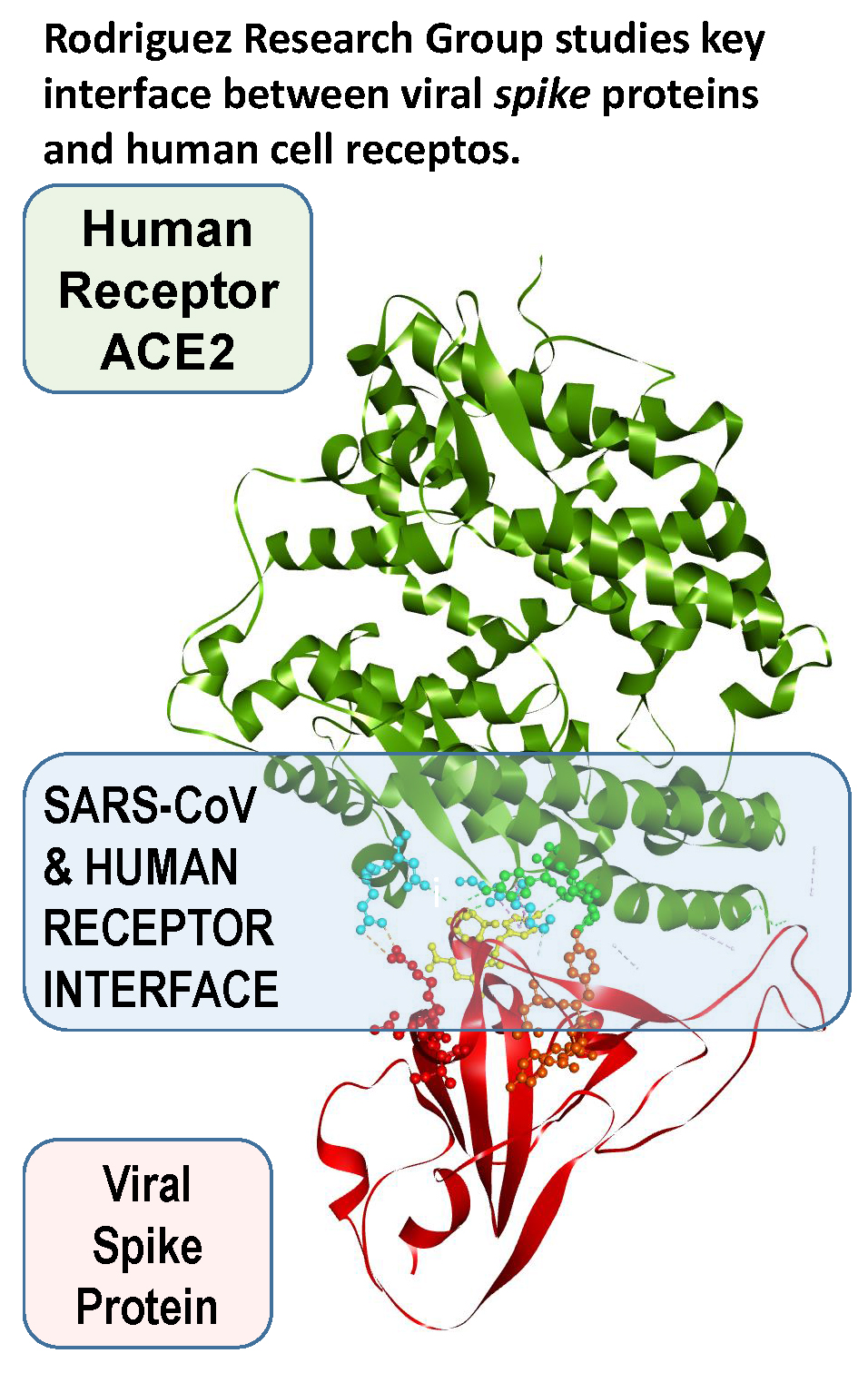
According to Rodríguez, “SARS-CoV-1 viruses interact with human ACE2 receptors (hACE2) via the receptor binding domains (RBD) of their spike (S) proteins. Structural characterization techniques such as X-ray crystallography (XRC) and cryogenic electron microscopy (cryo-EM) can identify the contact residues at the hACE2...S-RBM interface. However, these techniques cannot unequivocally determine which S-RBM residue fragments are attractive and which are repulsive relative to hACE2. By contrast, such information, helpful for antiviral or vaccine development, can be obtained via rigorous quantum-biochemical calculations as shown by recent work done in our laboratory. Our present computational studies are consistent with and, at the same time, complementary to those based on structural probes such as XRC and cryo-EM.”
Rodríguez and his group run a state-of-the-art high-performance computational lab at Purdue University which allows them to carry out specialized biomolecular simulations. The computational quantum-biochemical findings by the group may guide future efforts, carried out by pharmaceutical companies, to more efficiently develop antiviral drugs for coronaviruses. The research has identified specific viral-protein-sites which may be potentially targeted by new pharmaceuticals to combat potentially-reemerging SARS-CoV-1 viral infections.
This research is a rung in the ladder of SARS developments that Rodríguez and his team are working on. His lab is performing additional computational quantum-biochemical research on SARS-CoV-2, which is responsible for the recent outbreak of COVID-19 disease, and its mutations.
“We are now further studying, using advanced computational methods, the effects of recent mutations of the SARS-CoV-2 coronavirus (such as the N501Y mutation) recently discovered in the United Kingdom,” says Rodríguez. “These studies may help to explain why certain mutations are associated with an increased rate of infection in the UK population and, in addition, predict if current antiviral drugs or vaccines against SARS-CoV-2 (which is responsible for the current COVID-19 pandemic) may be effective or not.”
Additional information:
Prof. Jorge H. Rodríguez, E-mail: jhrodrig@purdue.edu
Computational (Bio)Molecular Physics Group, E-mail: bionanophys@purdue.edu